Programme
September 23rd ― Introduction and Phylogenetics
8.30-12.30
Gerd B. Müller(Konrad Lorenz Institute, Vienna)
Introduction to Evo-devo and the Extended synthesis in Evolutionary Theory
This unit will provide an overview of the conceptual background of evolutionary developmental biology, highlighting how new concepts, such as developmental constraint, facilitated variation, epigenetic innovation, or dynamical patterning, in concert with advances in other domains, contribute to an extended theoretical framework of evolutionary biology.
Ronald Jenner (Natural History Museum, London)
The role of phylogenetics in evo-devo
Developmental biology only becomes evo-devo by the addition of a phylogenetic context. First I will introduce the important roles that phylogenetic methods and data play, or should play, in current evo-devo research. Then I will address three questions that will highlight some of the greatest remaining challenges in phylogenetics, as well as draw attention to some of the inevitable limitations to our understanding even when we have a fully resolved tree in hand: 1) Will we ever be able to resolve the higher-level phylogeny of Metazoa?; 2) What is the predictive power of the phylogenetic position of taxa?; and 3) Evo-devo after the tree: can we bridge the gaps between extant body plans?
13.30-16.30
Phylogenetics Practical (Jenner)
16.30-dinner time
(Voluntary) Poster Session for Participants
September 24th ― Macroevolution and Morphology
8.30-12.30
Gonzalo Giribet (Harvard University)
Comparative Genomics/Body Plan Evolution
The origins and diversification of organisms is best understood by the study of body plan disparity and its evolution. Although traditionally studied from an anatomical perspective, the understanding of animal diversity has rapidly turned towards molecular information, especially with the increase in available sequence data and the rapid development of the techniques that allow researchers to obtain genome-level information. The emerging field of transcriptomics has allowed us to tackle questions ranging from phylogenetic patterns to ecological and developmental aspects of animals. But most important, the broader field of genomics has opened new doors into how we understand the evolution of body plan evolution. We will explore new research in animal genomics and its applications to the fields of evolution and development.
Graham Budd (University of Uppsala)
Constraining the Unconstrainable? Fossils and the Phenotype-Genotype Map
All evolutionary change took place in now extinct stem groups to living taxa and as a result must always remain partly shrouded in mystery. Given that one of the principal results of the field of evo-devo has been to emphasise the complexity of the genotype-phenotype map, it follows that even if we can recover the routes phenotypic evolution has taken, to unearth the true genotypic evolution that lay behind it will likely be highly problematic. This uncomfortable feature has led to a tendency in the literature to equate the (observed) results of genotypic evolution with the mechanism of the genotypic change driving (observed) phenotypic change – a move that I shall argue is invalid. If we are really to determine the patterns of causal evolution, we will need to know i) how far we can reconstruct the pattern of phenotypic change in the past (with the fossil record); ii) to what extent there is true freedom in the genotype-phenotype map within such phenotypic variation; and iii) if there are experimental ways of addressing what this freedom might consist of. I shall attempt to address each of these broad questions in turn without promising a solution to their most intractable aspects.
13.30-16.30
Journal Clubs (in small groups, moderated by teachers)
16.30-dinner time
Project Groups Assemble
September 25th ― (Post-)Transcriptional Regulation
8.30-12.30
Patricia Beldade (Gulbenkian Institute, Lisbon)
The Mechanisms for Variation and Diversity
Heritable phenotypic variation is the raw material of evolution by natural selection, and understanding its generation is a crucial issue in contemporary evolutionary biology. There is much current work investigating the mechanisms that underlie intra-specific variation and inter-specific diversity. I will use both classical examples and the very recent literature to provide an overview of how and which genes and environmental factors affect organismal development to account for different phenotypes. I will give special attention to three key and largely open questions in the field: What is the relationship between the genes responsible for intra-specific variation and inter-specific differences? What is the role of the external environment in producing phenotypic variation and how this matters for evolutionary change? How do novel traits arise and diversify?
Claudio Alonso (University of Sussex)
Post-transcriptional gene regulation during development and evolution
It is now indisputable that gene activity can be modulated through a wide spectrum of mechanisms, including transcriptional, epigenetic, as well as post-transcriptional processes such as microRNA regulation, RNA processing and post-translational modifications. Although the evolution of transcriptional regulation has attracted substantial attention in recent years, much less is known about the roles played by post-transcriptional regulation during the evolution of the gene regulatory programmes controlling animal development. To address this problem we shall first introduce a range of post-transcriptional mechanisms able to modify developmental gene function at the quantitative and/or qualitative level. We will then turn to discuss the ways by which variation in post-transcriptional regulation can be linked to the evolution of gene function. Building up on this background, we will subsequently explore a number of open questions within this emerging area of research, including: (i) How do transcriptional and post-transcriptional regulation relate to one another? (ii) How does microRNA expression relate to that of their gene targets? (iii) How do changes in mRNA expression level and distribution relate to modulations in protein level/distribution and, ultimately, how all this relates to the evolution of developmental function?
afternoon
Sightseeing in Venice
19.00-20.00
Alesssadro Minelli (University of Padova)
SPECIAL LECTURE: Morphological Misfits and the Architecture of Development
Misfits are a miscellaneous class of numerically marginal taxa, also possibly, but not necessarily, less successful in their adaptations than their ‘normal’ relatives. Morphological misfits offer a lot of potentially attractive case studies in evolutionary developmental biology. Anatomically, there are modular misfits such as paussine beetles, with extravagant antennae borne on a quite usual beetle body, and systemic misfits such as Sacculina. The former, but not the latter, are suggestive of developmental modularity. Ontogenetically, there are one phase-only misfits and whole life-cycle misfits. The former, but not the latter, would suggest an evolutionary independence of developmental stages. Phylogenetically, some misfits have diverged very recently from their morphologically conservative closest relatives, while others are the sole members of lineages that diverged very deeply in time from their closest relatives. The former, but not the latter, provide obvious opportunities for investigating character evolvability.
September 26th ― Networks and Physics
8.30-12.30
Johannes Jaeger (Center for Genomic Regulation – CRG, Barcelona)
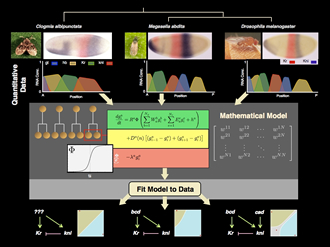
Evolving Networks/In Silico Evolution
I will give a brief overview on qualitative and quantitative approaches to the study of the dynamics, function, and evolution of developmental gene regulatory networks. We will then explore three central questions in evo-devo that can be addressed by network modelling: (1) how the structure and dynamics of gene regulatory networks affects the direction of evolutionary change through the production of non-random phenotypic variability; (2) how robustness and phenotypic plasticity enable and affect the evolvability of developmental systems; and finally (3) whether there are rules or regularities governing the evolution of organismic form, and how such rules could be uncovered.
Veronica Grieneisen (John Innes Centre, Norwich, UK)
Tissue Mechanics & Polarity in Animals and Plants
Firstly, we will discuss several categories of modelling formalisms that can capture properties of cell behaviour (such as cell shape) within tissues. We will discuss advantages and properties of these models, their strengths and limitations in face of specific biological systems. Special emphasis will be given on the Cellular Potts Model. We will move on and discuss the evo-devo aspects of cell polarity and link this to questions from how isolated cells spontaneously polarize up to how they coordinate their orientations within tissues. Here, many basic questions are still open in the field: how do tissues establish long-distance organization through local interactions? How to prevent tissue heterogeneities to interfere with such a coordination? How relevant is the mechanism of the cells' intrinsic polarity to the way it establishes coordination with neighbouring cells (and vice-versa)? I will present recent work elucidating the molecular mechanisms underlying these higher-level questions, by focusing on small G-protein interactions in both motile cells as well as growing plant cells. Interestingly, we will see that basic cell polarity mechanisms and their principles are extremely conserved over kingdoms, which provokes the next level of questions as to how multicellularity, that evolved independently in plants and animals, was guided by divergent mechanisms of tissue polarity coordination in animals, where direct cell-cell coupling is possible, and plant tissues, where only indirect cell-cell coupling is possible, due to the cell wall. Given that there are obvious differences, we will analyse to what level similarities within tissue polarity mechanisms can be found. To fully appreciate these questions and their implications, an overview of the key challenges in plant development will be provided through the course.
13.30-16.30
Demo/Practical on Networks and Modelling (Grieneisen/Jaeger)
September 27th ― Evolutionary Genetics and Ecology
8.30-12.30
Alistair McGregor (Oxford Brookes University)
Evo-Devo & Evolutionary Genetics
Micro-evo-devo represents the synthesis of evolutionary developmental biology and population genetics. It has long been recognized that by bringing a developmental and phenotypic focus to population genetics on one hand and population/quantitative genetic approaches to development and morphology on the other, micro-evo-devo has great potential to address how genotypic variation changes developmental programs to give rise to phenotypic differences as well as describe the evolutionary forces involved. However, the relative dearth of micro-evo-devo studies may risk pushing evo-devo to the periphery of the larger field of evolutionary developmental biology. Therefore, in my lectures I will restate the case for micro-evo-devo and then explore how micro-evo-devo approaches have helped to address three important questions vital to understanding the evolution and development of organisms: (1) What is the contribution of standing genetic variation to species differences? (2) What is the genetic basis of changes in complex quantitative traits? (3) What evolutionary forces have shaped phenotypic diversification?
Christen Mirth (Instituto Gulbenkian de Ciencia, Oeiras, Portugal)
Phenotypic Plasticity and the Evolution of Polyphenisms (Eco-Devo)
The environment moulds the development of virtually every organism. Sometimes differences in the rearing environment result in dramatically different phenotypes, or polyphenisms, from the same genotypes. A common example of this is nutritional differences giving rise to queen versus worker bees. Nevertheless, we understand very little about how these traits are regulated and how they evolve. Further, the role of phenotypic plasticity in evolution has been undervalued. In this module, we will explore three unanswered questions: (1) What are the mechanisms underlying phenotypic plasticity? (2) How do polyphenisms evolve? (3) How does plasticity contribute to the diversification of species?
13.30-16.30
Project Presentations
16.30-18.30
Closing Discussion |